Research project Source identification of brown rot and ring rot infections in Belgium (1989-2021) by forensic investigation of bacterial genome sequences
General introduction
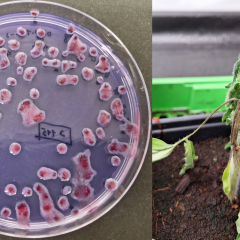
For more than 3 decades the quarantine bacteria brown rot (Ralstonia solanacearum) and ring rot (Clavibacter sepedonicus) have posed a danger to the cultivation of potatoes in particular, but can also infect tomato and eggplant. Still, farmers in some regions are not allowed to use surface water for irrigation of these crops due to the high risk of contamination. Between 1989 and 2021, samples were taken annually from waterways and from bittersweet, a host plant growing on riverbanks. Together with the bacteria isolated from infected batches of potatoes, all of these samples were kept at ILVO. In this research project, we use a reverse contact tracing approach with advanced DNA sequence analysis techniques. By trying to link all these samples together, we hope to trace the original source of infection. New sampling of waterways in the protection zone provides an up-to-date picture of the infection. This can then finally be used to reduce the number of municipalities in the protection zone and reduce the area in which use of surface water for potato cultivation is not allowed.
Research approach
Genetic analyses are performed on isolates of Ralstonia and Clavibacter obtained over the years as well as those kept in a culture collection. The basis for microbial genotyping in outbreak studies is Whole Genome Sequencing (WGS). The genomic data are then exploited through various forensic methods to map minute genetic differences. Some examples of these forensic techniques include MLVA-based typing (Multi Locus Variable Number Tandem Repeat Analysis), cg/ag MLST-based typing (core genome/accessory genome Multilocus Sequence Typing), SNP-based typing (Single Nucleotide Polymorphism)-based typing and CRISPR-based typing (Clustered Regularly Interspaced Short Palindromic Repeats).
Relevance/Valorization
This project will provide researchers with the necessary genetic information to accurately connect isolates from new infection clusters with each other and, if necessary, with previous infection(s). This should make it possible to reduce the number of protection zones in Belgium. The first results are expected by the end of 2024.